The Ghavi-Helm Laboratory
Enhancer-promoter interactions
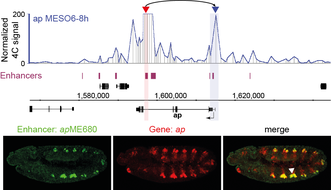
Our recent work suggests that (i) developmental genes are often regulated by multiple enhancers, sometimes located at great linear distances, (ii) the spatio-temporal activity of a large fraction of those enhancers remains unknown, (iii) enhancer-promoter interactions are usually established before the target gene is expressed and are largely stable during embryogenesis, and (iv) stable interactions seem to be associated with the presence of paused RNA Polymerase II at the promoter before gene activation.
Research plans
3D regulatory genomics during Drosophila embryogenesis
Building upon these results, we propose to advance to the next level in the dissection of enhancer-promoter interaction functionality in the context of Drosophila embryogenesis. We are investigating multiple questions, including but not limited to:
1) What determines the specificity of promoter-enhancer interactions in a complex genome?
2) Are enhancer-promoter interactions tissue-specific, and what are the drivers of this specificity?
3) Are all enhancer-promoter interactions functional, and how does the activity of an enhancer relate to the expression of the gene it interacts with?
Experimental techniques and equipment
The main tools used in the lab are:
4C/Hi-C/Capture-C, RNA-seq, ChIP-seq, ATAC-seq, CRISPR-Cas9 genome editing, FACS sorting, Drosophila genetics and transgenesis, bioinformatics.
Our favorite model organism is the fruit fly Drosophila melanogaster.
Thanks to generous funding from the ERC, we have recently acquired a SONY SH800 cell sorter:


We benefit from the core infrastructure available at the IGFL including shared facilities for animal housing, cell culture, microscopy, an in-situ hybridization robot, and a large particle sorter (Union Biometrica BioSorter) to sort intact embryos based on size and fluorescent pattern. The IGFL has an in-house NGS service with Illumina, Ion Proton and MinION Nanopore technologies. Moreover, we will have access to state-of-the-art computing clusters (>500 compute nodes) and use the Core Facilities of the Gerland Campus located in close proximity to the Institute.

credits: drosophiladrawings.blogspot.de
Funding







Follow us on twitter:
Institut de Génomique Fonctionnelle de Lyon (IGFL)
ENS de Lyon - CNRS UMR 5242 - INRA USC 1370
32-34 avenue Tony Garnier, 69007 Lyon, France



